Predict protein domains likely to mediate binding
Vitalii Kleshchevnikov
23/09/2018
protein_domains_binding.RmdLoad example dataset
These are interactions of viral and human proteins along with protein domain annotations. The aim of the analysis is to predict domains in human proteins likely to bind viral proteins. More details and be found in a paper.
data = fread("../data/viral_human_net_w_domains", sep = "\t", stringsAsFactors = F)library(igraph)
prot2domain = as_adjacency_matrix(graph.data.frame(unique(data[, .(IDs_interactor_human, IDs_domain_human)]))) prot2domain = prot2domain[rownames(prot2domain) %in% data\(IDs_interactor_human, colnames(prot2domain) %in% data\)IDs_domain_human]
viral2human = as_adjacency_matrix(graph.data.frame(unique(data[, .(IDs_interactor_viral, IDs_interactor_human)]))) viral2human = viral2human[rownames(viral2human) %in% data\(IDs_interactor_viral, colnames(viral2human) %in% data\)IDs_interactor_human]
viral2human %*% prot2domain
Find enriched domains
First, let’s use multi-core parallelisation. Correct analysis would require many more premutations but this is just an example of how to use this package.
res = permutationPval(interactions2permute = IDs_interactor_viral ~ IDs_interactor_human,
associations2test = IDs_interactor_viral ~ IDs_domain_human,
# node_attr gives a way to add columns needed to compute statistic and filter the data,
# in this case only domain_count is needed to filter
# and node columns to compute statistic
node_attr = list(IDs_interactor_viral ~ IDs_interactor_viral_degree,
IDs_domain_human ~ domain_count,
IDs_interactor_viral + IDs_domain_human ~ domain_frequency_per_IDs_interactor_viral),
data = data,
# in this example statistic is just count of
# IDs_interactor_human for each IDs_interactor_viral / IDs_domain_human pair
statistic = IDs_interactor_viral + IDs_domain_human ~ .N,
select_nodes = IDs_domain_human ~ domain_count >= 1,
N = 1, # number of permutations
cores = 1, # how many cores to use on a local machine
# computations are split into inner and outer replicate
# to help manage memory load, here it is 50*2
# hint: you can use microbenchmark package to find optimal value
split_comp_inner_N = 1,
seed = 2) # seed for reproducible sampling (permutations)Next, let’s try multi-node parallelisation (on a computing cluster). Default installation of clustermq will still work but run multiple processes on a local machine.
res = permutationPval(interactions2permute = IDs_interactor_viral ~ IDs_interactor_human,
associations2test = IDs_interactor_viral ~ IDs_domain_human,
node_attr = list(IDs_domain_human ~ domain_count),
data = data,
statistic = IDs_interactor_viral + IDs_domain_human ~ .N,
select_nodes = IDs_domain_human ~ domain_count >= 1,
N = 30,
clustermq = T, # use clustermq
clustermq_jobs = 3, # how many cluster jobs to start
# how much memory each job needs
clustermq_mem = 2000,
# caution: allocating more memory than needed is a waste of resources,
# but allocating not enough will result in you jobs being killed,
# in turn, this will freese R requiring restart
# hint: try 2-3 jobs with realistic load and excess of memory
split_comp_inner_N = 2, seed = 2)## Submitting 3 worker jobs (ID: 7716) ...Look at the results and p-value distribution
res##
## This object contains the empirical p-value for the association between <IDs_interactor_viral> and <IDs_domain_human> both of which are linked (connected through edges) by <IDs_interactor_human>
##
## Produced by permutationPval function call:
##
## permutationPval(interactions2permute = IDs_interactor_viral ~
## IDs_interactor_human, associations2test = IDs_interactor_viral ~
## IDs_domain_human, node_attr = list(IDs_domain_human ~ domain_count),
## data = data, statistic = IDs_interactor_viral + IDs_domain_human ~
## .N, select_nodes = IDs_domain_human ~ domain_count >=
## 1, N = 30, seed = 2, clustermq = T, clustermq_mem = 2000,
## clustermq_jobs = 3, split_comp_inner_N = 2)
##
## $data_with_pval is the data.table containing the original data and appended with empirical p-value (p.value) as well as observed_statistic, YmissingZ_perX, and higher_counts, not_missing used to calculate p-value,
## Details: `?permutationPval`
##
## x$data_with_pval
## IDs_interactor_viral IDs_domain_human observed_statistic
## 1: P03427 IPR001909 6
## 2: P03427 IPR013083 15
## 3: P03427 IPR013087 9
## 4: P03427 IPR013087 9
## 5: P03427 IPR011650 1
## ---
## 45518: P04591 IPR015915 1
## 45519: P04591 IPR015916 1
## 45520: Q77ZG4 IPR001660 1
## 45521: Q77ZG4 IPR004177 1
## 45522: Q77ZG4 IPR013761 1
## YmissingZ_perX higher_counts not_missing p.value
## 1: 18 966 55230 0.0174904943
## 2: 18 75 138075 0.0005431831
## 3: 18 414 82845 0.0049972841
## 4: 18 414 82845 0.0049972841
## 5: 18 9205 9205 1.0000000000
## ---
## 45518: 1 856 856 1.0000000000
## 45519: 1 856 856 1.0000000000
## 45520: 1 413 413 1.0000000000
## 45521: 1 413 413 1.0000000000
## 45522: 1 413 413 1.0000000000
## IDs_interactor_human
## 1: A0A024R0Q3
## 2: A0A024R0Q3
## 3: A0A024R0Q3
## 4: A0A024R0Q3
## 5: A0A024R382
## ---
## 45518: Q9Y6Y0
## 45519: Q9Y6Y0
## 45520: Q9Y6Y8
## 45521: Q9Y6Y8
## 45522: Q9Y6Y8
## all_IDs_domain
## 1: PS50805|PF01352|SSF109640|SM00349|cd07765
## 2: G3DSA:3.30.40.10
## 3: PS00028|PS00028|SSF57667|PS50157|PS00028|PS00028|PS50157|SSF57667|PS50157|PS50157|SSF57667|PS50157|PS00028|PS00028|SSF57667|SSF57667|SSF57667|PS00028|PS00028|PS50157|SM00355|SM00355|SM00355|SM00355|SM00355|SM00355|SM00355|SM00355|SM00355|SM00355|SM00355|SM00355|SM00355|SM00355|SM00355|SM00355|PS00028|PS00028|PS50157|PS50157|PS50157|PS00028|PS00028|PS50157|PS50157|SSF57667|SSF57667|PS00028|PS00028|PS50157|SSF57667|PS50157|PS00028|PS00028|PS50157|PS50157|PS50157
## 4: PS00028|SM00355|SM00355|PS50157|SSF57667|PS50157|PS00028
## 5: PF07687
## ---
## 45518: G3DSA:2.120.10.80
## 45519: G3DSA:2.130.10.80
## 45520: SM00454|PF00536
## 45521: PF02862|SM01127
## 45522: SSF47769
## domain_type N_prot_w_interactors domain_count domain_frequency
## 1: Domain 15940 278 0.0174404015
## 2: Domain 15940 862 0.0540777917
## 3: Domain 15940 712 0.0446675031
## 4: Domain 15940 712 0.0446675031
## 5: Domain 15940 6 0.0003764115
## ---
## 45518: Domain 15940 73 0.0045796738
## 45519: Domain 15940 21 0.0013174404
## 45520: Domain 15940 111 0.0069636136
## 45521: Domain 15940 8 0.0005018821
## 45522: Domain 15940 140 0.0087829360
## Taxid_interactor_human Taxid_interactor_viral
## 1: 9606 381518
## 2: 9606 381518
## 3: 9606 381518
## 4: 9606 381518
## 5: 9606 381518
## ---
## 45518: 9606 11706
## 45519: 9606 11706
## 45520: 9606 37296
## 45521: 9606 37296
## 45522: 9606 37296
## IDs_interactor_viral_degree IDs_interactor_human_degree
## 1: 168 1
## 2: 168 1
## 3: 168 1
## 4: 168 1
## 5: 168 1
## ---
## 45518: 11 1
## 45519: 11 1
## 45520: 5 1
## 45521: 5 1
## 45522: 5 1
## IDs_domain_human_per_IDs_interactor_viral
## 1: 280
## 2: 280
## 3: 280
## 4: 280
## 5: 280
## ---
## 45518: 37
## 45519: 37
## 45520: 13
## 45521: 13
## 45522: 13
## IDs_interactor_viral_per_IDs_domain_human
## 1: 25
## 2: 201
## 3: 108
## 4: 108
## 5: 2
## ---
## 45518: 21
## 45519: 17
## 45520: 31
## 45521: 3
## 45522: 44
## domain_count_per_IDs_interactor_viral
## 1: 6
## 2: 15
## 3: 9
## 4: 9
## 5: 1
## ---
## 45518: 1
## 45519: 1
## 45520: 1
## 45521: 1
## 45522: 1
## domain_frequency_per_IDs_interactor_viral fold_enrichment
## 1: 0.035714286 2.047790
## 2: 0.089285714 1.651061
## 3: 0.053571429 1.199338
## 4: 0.053571429 1.199338
## 5: 0.005952381 15.813492
## ---
## 45518: 0.090909091 19.850560
## 45519: 0.090909091 69.004329
## 45520: 0.200000000 28.720721
## 45521: 0.200000000 398.500000
## 45522: 0.200000000 22.771429plot(res)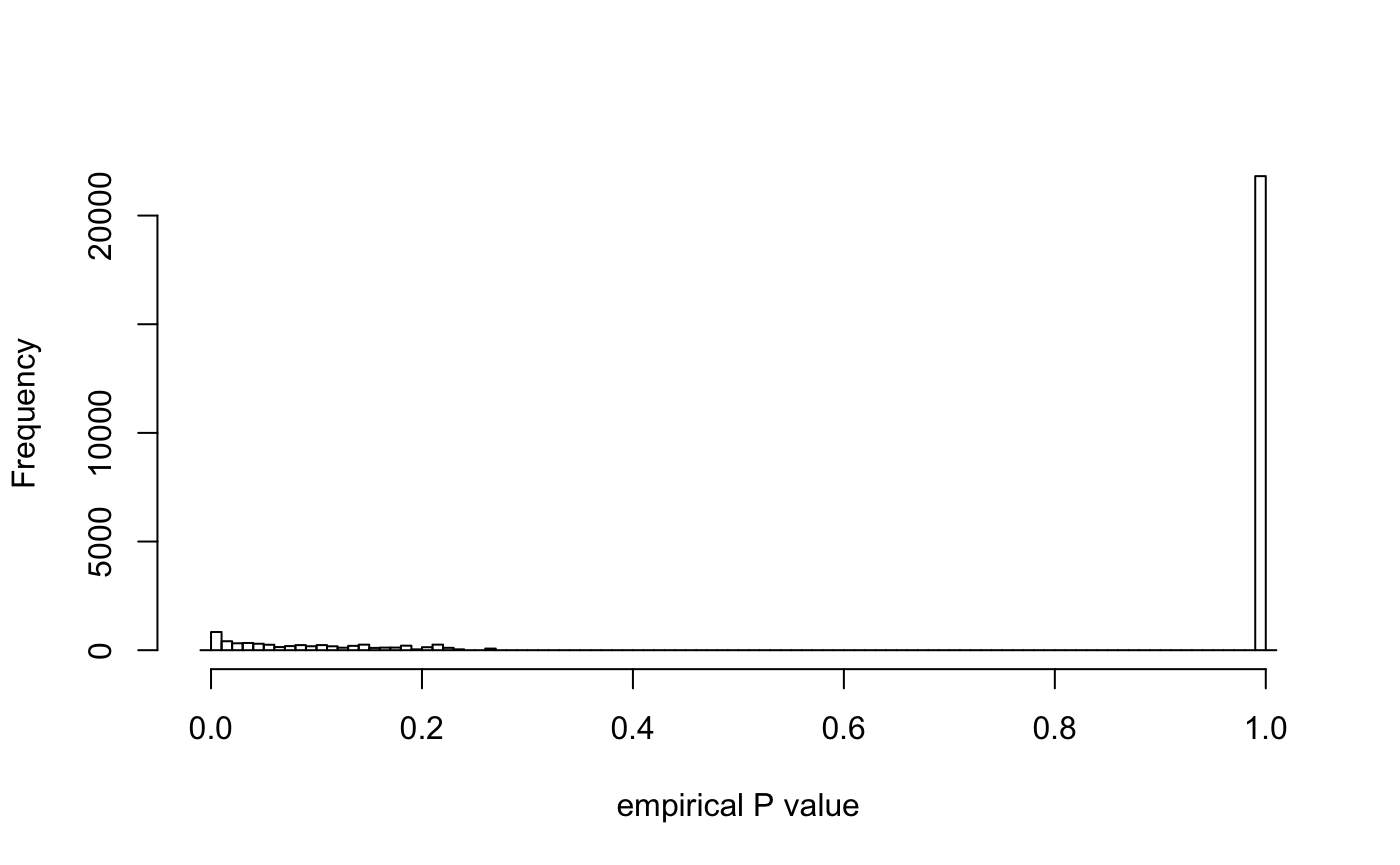
Date and packages used
Sys.Date. = Sys.Date()
Sys.Date.## [1] "2018-11-20"session_info. = devtools::session_info()
session_info.## ─ Session info ──────────────────────────────────────────────────────────
## setting value
## version R version 3.5.1 (2018-07-02)
## os macOS High Sierra 10.13.6
## system x86_64, darwin15.6.0
## ui X11
## language (EN)
## collate en_GB.UTF-8
## ctype en_GB.UTF-8
## tz Europe/London
## date 2018-11-20
##
## ─ Packages ──────────────────────────────────────────────────────────────
## package * version date lib
## assertthat 0.2.0 2017-04-11 [1]
## backports 1.1.2 2017-12-13 [1]
## base64enc 0.1-3 2015-07-28 [1]
## bindr 0.1.1 2018-03-13 [1]
## bindrcpp 0.2.2 2018-03-29 [1]
## callr 3.0.0 2018-08-24 [1]
## cli 1.0.1 2018-09-25 [1]
## clustermq 0.8.5 2018-09-29 [1]
## colorspace 1.3-2 2016-12-14 [1]
## commonmark 1.6 2018-09-30 [1]
## crayon 1.3.4 2017-09-16 [1]
## data.table * 1.11.8 2018-09-30 [1]
## desc 1.2.0 2018-05-01 [1]
## devtools 2.0.1 2018-10-26 [1]
## digest 0.6.18 2018-10-10 [1]
## dplyr 0.7.8 2018-11-10 [1]
## evaluate 0.12 2018-10-09 [1]
## fs 1.2.6 2018-08-23 [1]
## GGally 1.4.0 2018-05-17 [1]
## ggplot2 3.1.0 2018-10-25 [1]
## glue 1.3.0 2018-07-17 [1]
## gtable 0.2.0 2016-02-26 [1]
## hms 0.4.2 2018-03-10 [1]
## htmltools 0.3.6 2017-04-28 [1]
## knitr 1.20 2018-02-20 [1]
## lazyeval 0.2.1 2017-10-29 [1]
## magrittr 1.5 2014-11-22 [1]
## MASS 7.3-50 2018-04-30 [2]
## memoise 1.1.0 2017-04-21 [1]
## munsell 0.5.0 2018-06-12 [1]
## NetFeaturePval * 0.1.0 2018-11-20 [1]
## pillar 1.3.0 2018-07-14 [1]
## pkgbuild 1.0.2 2018-10-16 [1]
## pkgconfig 2.0.2 2018-08-16 [1]
## pkgdown 1.0.0 2018-05-03 [1]
## pkgload 1.0.2 2018-10-29 [1]
## plyr 1.8.4 2016-06-08 [1]
## prettyunits 1.0.2 2015-07-13 [1]
## processx 3.2.0 2018-08-16 [1]
## progress 1.2.0 2018-06-14 [1]
## ps 1.2.1 2018-11-06 [1]
## purrr 0.2.5 2018-05-29 [1]
## qvalue 2.14.0 2018-10-30 [1]
## R6 2.3.0 2018-10-04 [1]
## RColorBrewer 1.1-2 2014-12-07 [1]
## Rcpp 1.0.0 2018-11-07 [1]
## remotes 2.0.2 2018-10-30 [1]
## reshape 0.8.8 2018-10-23 [1]
## reshape2 1.4.3 2017-12-11 [1]
## rlang 0.3.0.1 2018-10-25 [1]
## rmarkdown 1.10 2018-06-11 [1]
## roxygen2 6.1.1 2018-11-07 [1]
## rprojroot 1.3-2 2018-01-03 [1]
## rstudioapi 0.8 2018-10-02 [1]
## rzmq 0.9.4 2018-09-18 [1]
## scales 1.0.0 2018-08-09 [1]
## sessioninfo 1.1.1 2018-11-05 [1]
## stringi 1.2.4 2018-07-20 [1]
## stringr 1.3.1 2018-05-10 [1]
## testthat 2.0.1 2018-10-13 [1]
## tibble 1.4.2 2018-01-22 [1]
## tidyselect 0.2.5 2018-10-11 [1]
## usethis 1.4.0 2018-08-14 [1]
## withr 2.1.2 2018-03-15 [1]
## xml2 1.2.0 2018-01-24 [1]
## yaml 2.2.0 2018-07-25 [1]
## source
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.1)
## CRAN (R 3.5.0)
## CRAN (R 3.5.1)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.1)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.1)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## Github (vitkl/NetFeaturePval@cac7088)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.1)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## Bioconductor
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.1)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.1)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.1)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
## CRAN (R 3.5.0)
##
## [1] /Users/vk7/Library/R/3.5/library
## [2] /Library/Frameworks/R.framework/Versions/3.5/Resources/library